Leaflet identifyer
- Details
- Last Updated: Wednesday, 06 October 2021 07:56

A brand new tool to detect topological changes in lipid systems, as part of the PhD work of Bart Bruininks and Albert Thie: pubs.acs.org/doi/abs/10.102
 General Purpose Coarse-Grained Force Field
General Purpose Coarse-Grained Force Field 


























































































































































A brand new tool to detect topological changes in lipid systems, as part of the PhD work of Bart Bruininks and Albert Thie: pubs.acs.org/doi/abs/10.102
As part of the recent online Martini workshop, we developed a large number of new tutorials featuring the latest Martini 3 models. You can find the tutorials here, as well as a number of webinars on various aspects of Martini 3 and related tools. Enjoy !
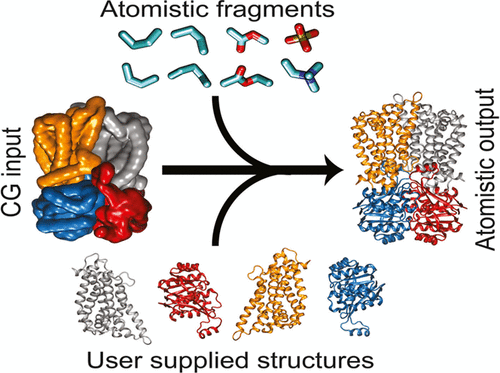
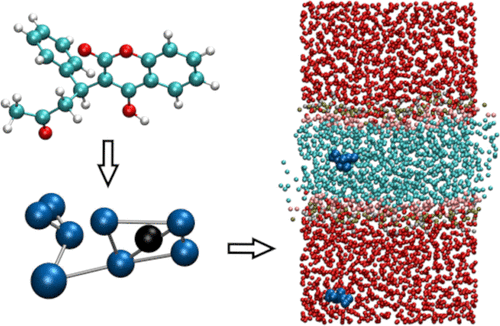 New Martini tools are popping up like snails in my backyard, with the difference that the tools are welcome additions whereas the snails are not.
New Martini tools are popping up like snails in my backyard, with the difference that the tools are welcome additions whereas the snails are not.
Check these papers for an updated backmapping tool from the Stansfeld group, and a brand new ATB from Miller and coworkers.

The Martini workshop was a great success - thanks to all who contributed to the meeting. The gather town setting made us feel like we were actually interacting with each other, and the cocktails at the bar almost tasted real. We are already thinking about another such meeting during winter time, but of course hope to welcome you in Groningen for an onsite meeting in due time !
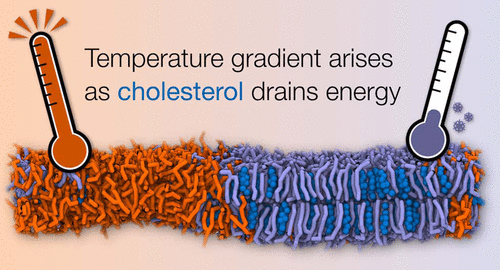
Be careful simulating membranes containing cholesterol to avoid temperature gradients developing. Read how to handle these systems in the publication that just appeared in JPCB, by Thallmair et al.: https://pubs.acs.org/doi/full/10.1021/acs.jpcb.1c03665

Together with researchers from Utrecht University, we have been able to visualize the 'molecular postman' that delivers important proteins, like antibodies and some hormones, to the outside of the cell. The research is published today in Molecular Cell: https://authors.elsevier.com/